For many years, Dr. Cothran did the testing at UKY.,
(Equine Blood Typing Research Laboratory Department of Veterinary Science)
Now Dr. Cothran is furthering his work at Texas A&M.
Federal law mandates that wild horses on public lands be managed in a manner that preserves both the horses and the habitat. The achievement of such a balance is not a simple matter and preservation of the habitat frequently requires that the horse population be kept small. However, small population size is a major risk factor for populations both in terms of susceptibility to catastrophes and genetics. From a genetic standpoint small population size can lead to the loss of genetic variation and inbreeding. Loss of genetic variation can threaten long term adaptability while inbreeding can result in reduced viability or fertility.
It is possible to manage populations so that the effects of small population size upon genetic variability can be minimized. An important first step to developing management strategies is an assessment of current levels of genetic variation within the population. In this report I present the results of genetic marker analysis of two feral horse populations managed by the Cedar City District of the LM in Utah. These are the Sulphur Herd Management Area (HMA) population and the Chloride Canyon HMA population.
Materials and Methods
A total of 118 horses from the Sulphur HMA were collected and analyzed. Most of these samples were provided by the BLM, however, some were obtained from horses that had been adopted. All were wild born animals. For both herds the majority of samples were collected in 1995. Blood samples were sent to the University of Kentucky for analysis. The samples were separated into red blood cells (rbd), rbc lysate, and serum. Standard immunological procedures involving hemagglutination and complement mediated hemolysis (Stormont and Suzuki, 1964; Stormone et al.,1964) were used to detect variation of red cell alloantigens at seven blood group loci. Starch and polyacrylimide gel electrophoresis and isoelectric focusing were used to detect variation at 10 serum and rbc lysate protein loci (Braend, 1973; Sandberg, 1974; Juenta et al., 1978; Braend and Johansen, 1983; Pollitt and Bell, 1980; Henney et al., 1994).
The loci examined were the A, C, D, K, P, Q and U horse blood group (HBG) loci and the biochemical protein loci were alpha-1-beta glycoprotein (A1B), albumin (Al), serum esterase (Est), vitamin D binding protein (Gc), glucosephosphate isomerase (GPI), alpha-hemoglobin (Hb), 6-phosphogluconate dehydrogenase (6-PGD), phosphoglucomutase (PGM), protease inhibitor (Pi) and transferrin (Trf) loci. Nomenclature for variants at all 17 loci was in accord with internationally standardized usage for horses (Bowling and Clark, 1985; Bowling and Ryder, 1987) except for variants at some loci which have not yet received international recognition.
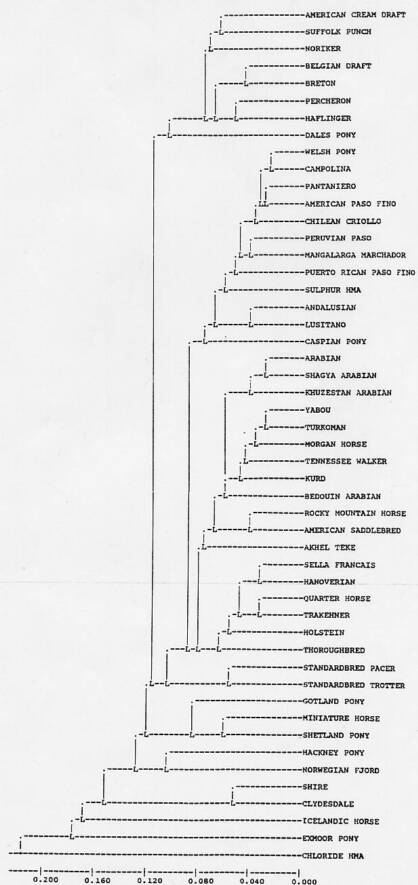
Gene frequencies for biochemical loci were calculated by direct count. Frequencies of alleles at blood group loci were calculated by the allocation method (Andersson, 1985). Genetic variation was measured as observed heterozygosity (Ho), Hardy-Weinberg expected heterozygosity (He), unbiased expected heterozygosity (Hu; Nei, 1978), effective number of alleles (Ae), and the total number of variants found in each population (Na). Ho was calculated for biochemical loci only because of the presence of recessive alleles and/or ambiguous genotypes at blood group loci. Therefore, for direct comparison, He and Hu were calculated for biochemical loci only (in an ideal population He (Hu) should equal Ho), for blood group loci, and for all 17 loci. In addition, population inbreeding level was estimated by Write's Fis=1-(Ho/He). Values of genetic variation of the Sulphur and Chloride herds were compared to those of 99 domestic horse populations that have been tested at the University of Kentucky. Genetic relationship of the Sulphur and Chloride horses to these other domestic breeds were investigated using Rogers' (1972) genetic similarity coefficient (S) and Nei's modified genetic distance (Da; Nei,et al., 1983) Restricted maximum likelihood analysis (RML Felsentein, 1989) and the UPGMA method were used to construct the dendrograms of Figures 1 and 2.
Results
Gene frequencies for both populations are given in Table 1. Measures of genetic variation for the Chloride and Sulphur herds are given in Table 2 along with values for breeds of domestic horses selected to show the range of variation of these measures in horse populations. Mean values for genetic variation measures for domestic and feral horses also are given in Table 2.
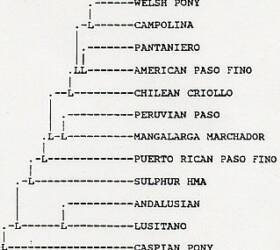
Measures of genetic similarity and distance of the Sulphur and Chloride Canyon herds to 73 breeds of horses are shown in Tables 3 and 4, respectively. Figure 1 shows estimated relationship of the Chloride Canyon and Sulphur horses to 50 breeds based upon RML analysis. The tree shown is a majority rule, strict consensus tree derived from 30 RML trees. The numbers at each node of the tree indicate how many times the arrangement to the right of the node occurred out of the 30 trees. Figure 2 shows estimated relationship of the Utah populations to domestic breeds based upon UPGMA cluster analysis of Nei's (1972) genetic distance.
Discussion
Levels of genetic variation in the Chloride and Sulphur feral herds were strikingly different (Table 2). Ho of the Sulphur herd was equal to average Ho for domestic horse breeds and higher than average for feral horses while He was well above the mean for both domestic and feral populations.
For the Sulphur herd, the Ho value, although about average for horses, indicates relatively good individual genetic health of the herd. The populational genetic diversity of the Sulphur herd (He,Ae and Na) was high compared to domestic and feral mean values. (Table 2). The Fis value for the Sulphur herd was positive. Although the Fis was not statistically significant, a positive Fis is suggestive of inbreeding. However, this probably is not the case for the Sulphur herd. The Sulphur population is divided into several different units which are somewhat distinct. I have analyzed some of these units individually (data not reported here due to uncertainty of the correct assignment of individuals to units) and within units the Ho is consistently greater than the He. Combining distinct subpopulations will normally result in He that is higher than Ho (a principal called the Whalund effect). The variability results show that there s a subdivision of the Sulphur herd and that overall, the variability of the Sulphur population is at a healthy level.
The second aspect of the genetic analysis of the Sulphur and Chloride herds was to determine the possible ancestral origins of these populations. Genetic distance and similarity analysis of the Sulphur herd to selected domestic breeds shows greatest resemblance to breeds of Iberian descent (Table3). The second closest mean similarity and distance was with a group of horses I call the gaited North American breeds. These breeds all have Spanish ancestry.
Highest individual similarity values were to the Chilean Criollo, Puerto Rican Paso Fino and American Paso Fino, all Spanish breeds. Mean similarity and distance to other major groups of breeds for the Sulphur herd were consistently lower. In addition to the genetic distance and similarity analysis, the Sulphur herd shows genetic variants that probably are of Iberian origins at the Trf, Est, Pi, A HBG, D HGB and Q HGB loci. In particular the D cfgkm and Q ac variants likely represent old Spanish blood lines. Both these alleles were at very low frequency, but their presence is significant.
The above data show fairly strong evidence that the Sulphur herd has Spanish ancestry. It is not possible to quantify this ancestry. Additionally, there is evidence that this population is not of pure Spanish ancestry, although again it is not possible to quantify the non-Spanish component of this herd. There is also one variant in the Est system that may be a mutation that occurred in the population since it has been wild. We have listed this variant as Est-G* in Table 1. We have never observed this variant in any other non-Sulphur horse and we have tested over 140 other populations and over 200,000 individuals.
The cluster analysis of the Sulphur herd with other selected breeds using the RML analysis does not appear to support the Spanish ancestry (Figure 1) However, this figure must not be taken at face value. The RML procedure is designed to determine ancestor descent relationships based upon genetic distance among populations within the analysis. An outgroup population (in this case the Przewalski horse) is used to determine the most ancient split among all the populations. In the case of domestic (and feral) populations or breeds, it is not possible to depict true ancestor-descendent relationships due to high degree of crossing among horse breeds throughout the history of domestic horses. However, it can produce dendrograms that show realistic associations among the breed.